-
Notifications
You must be signed in to change notification settings - Fork 2
Metaproteomics Data
First coined in 2004, Metaproteomics deals with identifying & quantifying proteins from microbial communities to study and experiment with features and behaviour of microorganisms at a molecular level. The data, we intend to deal with from our huge Clinical Knowledge Graph, are information on peptides, proteins & metaproteins to find a connection between their abundance patients with various diseases. Hence we put forward a summary of our keywords explained briefly & the connections they share to make our queries in Neo4j more feasible to understand.
Proteins, composed of amino acids, are the most essential building block of all living organisms on the planet & are directly involved in the chemical processes, essential for life. The word itself has been derived from the Greek word "prōteios", which means "holding first place". As Encyclopaedia Britannica puts it, proteins are
- species-specific (differs from one species to the other) as well as
- organ-specific i.e. for e.g. within a single organism, proteins in brain will be different to proteins in liver & so forth.
Note - Proteins of similar function, have similar aminino acid composition & sequence
Proteins & peptides are kind of similar through structural aspects. Peptides act as a subset for a protein consisting of between 2 - 50 amino acids held together by peptide bonds, whereas proteins consist of 50 or more amino acids
Metaproteins are protein groups that consider the special use case of metaproteomics. In order to deal with homologous proteins, which are expected in a multi-species system, proteins are grouped into metaproteins using a set of rules. The metaprotein will then be assigned to a taxonomy based on the protein it contains and also depending on the specification provided by a user. Unlike protein groups used by other proteomics tools, metaproteins should not be considered a single protein with an ambiguous identification, but instead, they constitute a group of related proteins all of which are potentially contained in the sample. From this it follows that metaproteins will sometimes be assigned apparently unspecific taxonomies (i.e. Superkingdom rank), which indicates that the protein sequences on which the metaprotein is based are highly conserved across different taxa, making a specific taxonomic assignment impossible in a microbial community of multiple unknown species. Metaproteins will also combine other metadata from its proteins into a single entry: UniProt Keywords, UniRef Clusters, KEGG Orthology and enzyme commission numbers (EC).
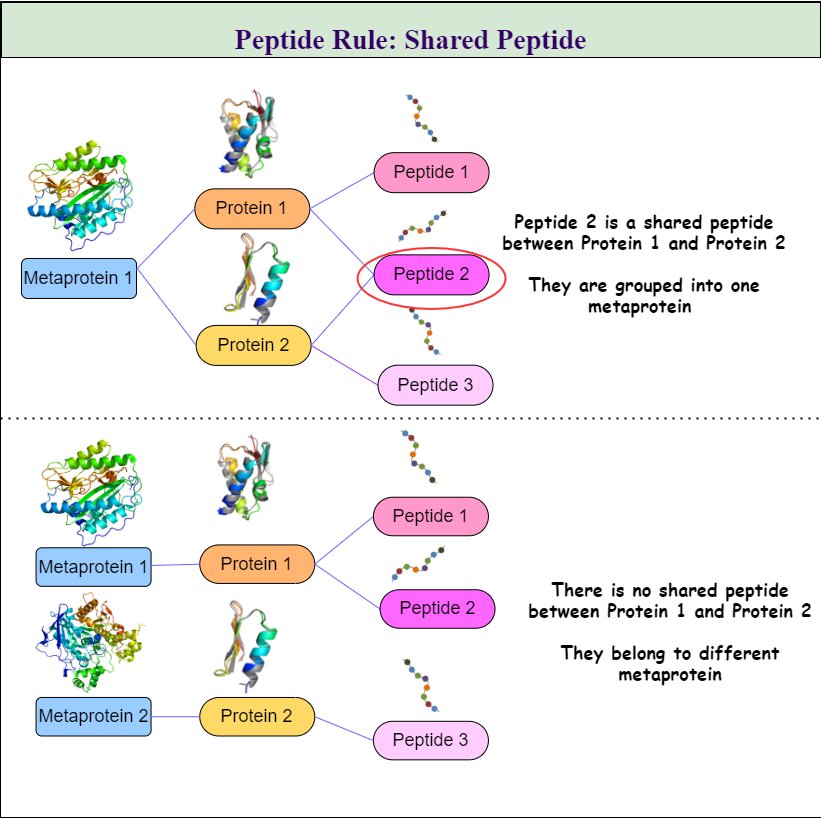
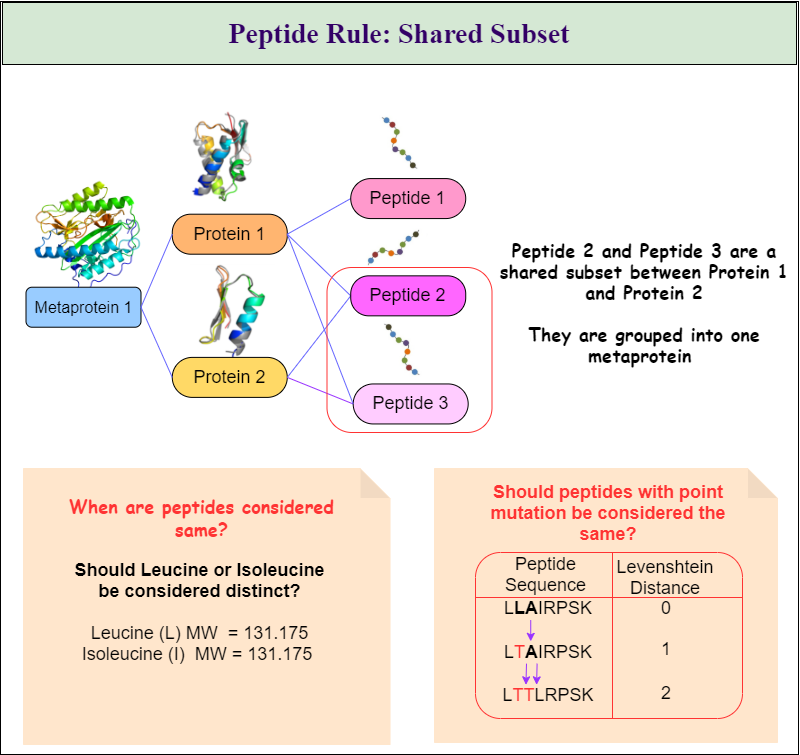
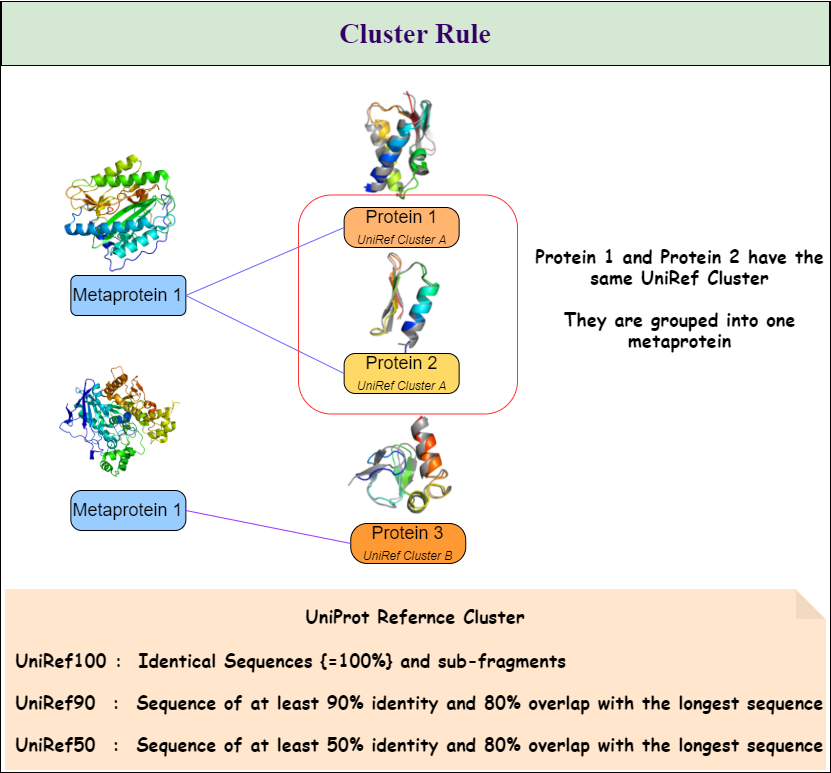
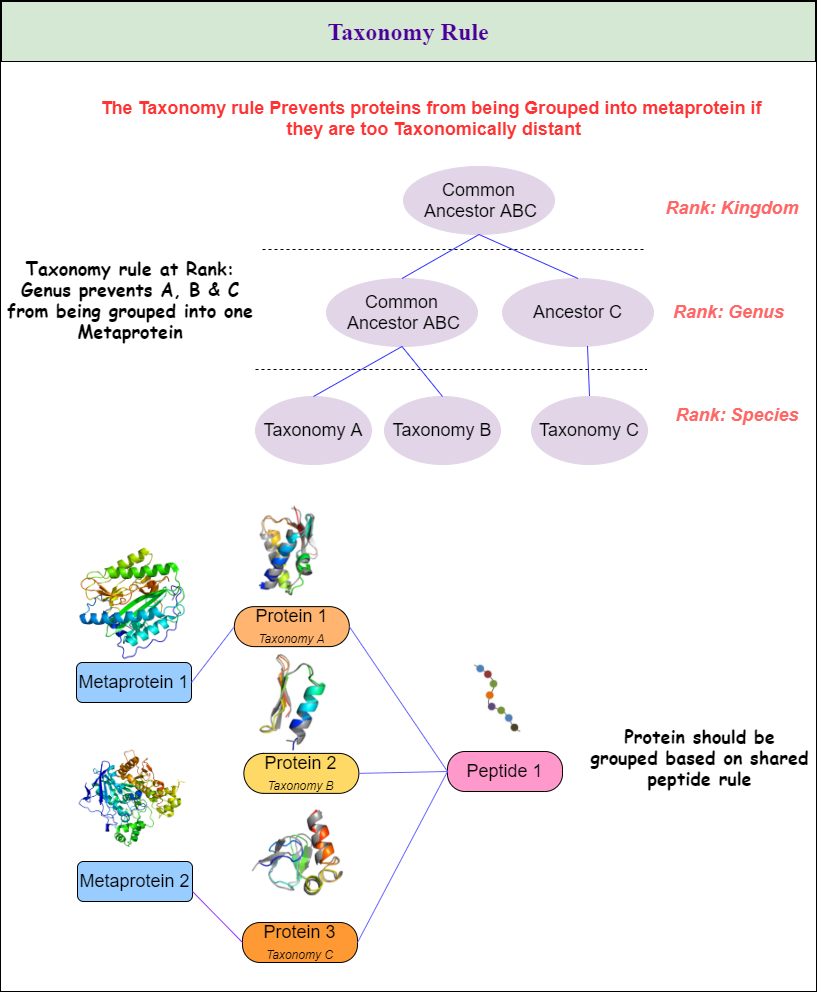
Metaproteins will be created according to the rules the user chooses. All three rules can be combined in any combination. The three rules are:
- Peptide Rule
- Cluster Rule and
- Taxonomy Rule as seen in Figure 1. \ Table 1 shows all available options and gives a description of how they will affect the metaprotein generation.